Oral squamous cell carcinomas (OSCC) often develops from oral potentially malignant disorders (OPMDs). Mostly due to late diagnosis, the mortality rate of OSSC has been at 50% for decades. The current standard of care for predicting progression from OPMDs consists of dysplasia grading, which shows highly variable behavior across patients and does not relate very well to the risk of developing cancer from the lesion.
Why Visiopharm’s Discovery software helps
Proteocyte used Visiopharm’s software to develop an APP that detects and quantifies S100A7 in the oral lesion sample.
The Stevenson-Lerner/Saldarriaga Lab team is involved in several translational research projects around chronic liver diseases, such as steatotic liver disease, chronic hepatitis C, autoimmune hepatitis, primary biliary cholangitis, and infectious diseases like Ebola. They investigate hepatic immunology, including macrophages and lymphocytes and how dysregulation of the immune response leads to fibrosis development as well as how the phenotype of a person’s hepatic macrophages determines downstream immune activation and how they respond to liver injury.
Why Visiopharm’s Discovery software helps
The team chose Visiopharm’s software for its ability to integrate complex biological data into a spatially resolved image analysis framework.
Autosomal dominant polycystic kidney disease (ADPKD) is an inherited systemic disorder mainly associated with mutations in the PKD1 gene and characterized by the development of multiple cysts in the kidneys and other organs. There is still a high unmet need for treatment options for patients with ADPKD because Tolvaptan, the only approved treatment, has limited efficacy and non-negligible side effects. To mimic naturally occurring human PKD1 mutations, a PKD1 inducible knockout mice strain was established at Novalix to study disease progression and evaluate therapeutic approaches.
Why Visiopharm’s Discovery software helps
The software enabled comprehensive histological and imaging analyses, providing actionable insights into the disease mechanisms and progression.
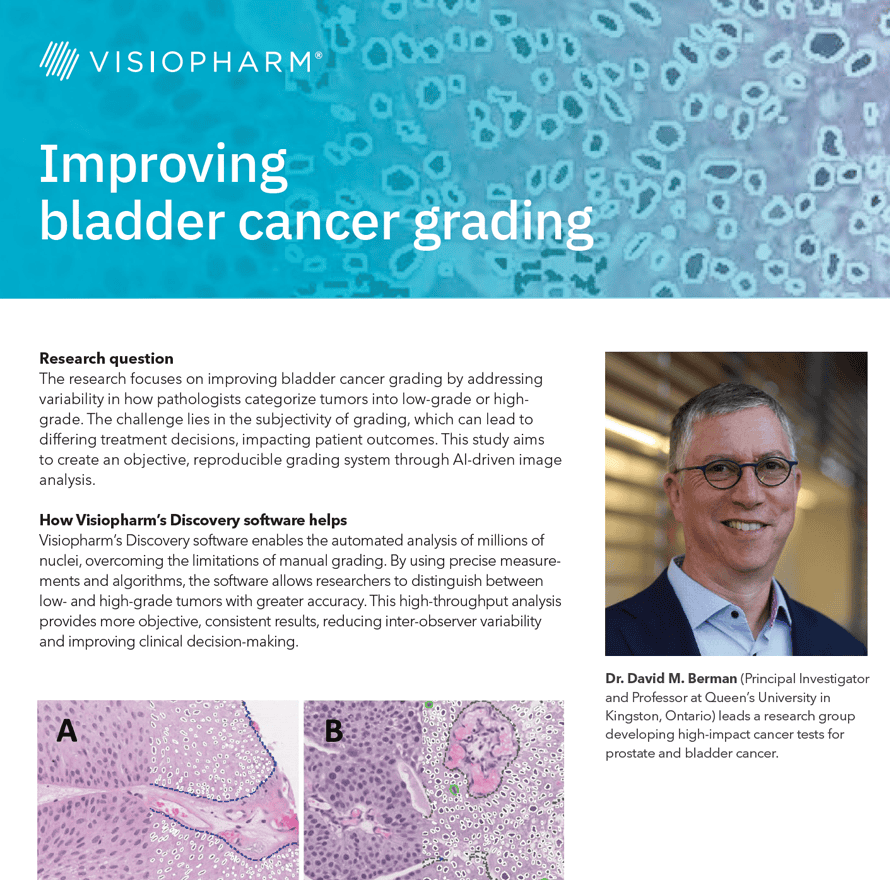
The research focuses on improving bladder cancer grading by addressing variability in how pathologists categorize tumors into low-grade or highgrade. The challenge lies in the subjectivity of grading, which can lead to differing treatment decisions, impacting patient outcomes. This study aims to create an objective, reproducible grading system through AI-driven image analysis.
How Visiopharm’s Discovery software helps
Visiopharm’s Discovery software enables the automated analysis of millions of nuclei, overcoming the limitations of manual grading.
The research focuses on improving bladder cancer grading by addressing variability in how pathologists categorize tumors into low-grade or highgrade. The challenge lies in the subjectivity of grading, which can lead to differing treatment decisions, impacting patient outcomes. This study aims to create an objective, reproducible grading system through AI-driven image analysis.
How Visiopharm’s Discovery software helps
Visiopharm’s Discovery software enables the automated analysis of millions of nuclei, overcoming the limitations of manual grading.
The high mortality rate of COVID-19 is largely due to acute respiratory distress syndrome (ARDS), which is characterized by diffuse alveolar damage (DAD). Additionally, severe cases of COVID-19 often result in a cytokine storm and a disrupted adaptive immune response. Most previous studies on this issue have focused on the peripheral cell count and the functionality of immune cells.
The team at UK Augsburg in Germany used multiplexed immunofluorescence to study the impact of SARS-CoV-2 on antigen-presenting cells. Like MERS-CoV and SARS-CoV, SARS-CoV-2 appears to impair the maturation of dendritic cells (DCs), which is characterized by a switch in surface antigen expression. This switch enables the cells to travel to lymph nodes and activate T-cells.
To shed light on the local inflammatory infiltrate, the team compared the cell populations of professional antigen-presenting cells (APCs) in the lungs of COVID-19 autopsy cases in various stages of DAD. They found an increased number of myeloid dendritic cells (mDCs) in later stages, but with no significant upregulation of maturation markers in DAD specimens with high viral load. This accumulation of immature mDCs, which are unable to reach lymph nodes, results in an inadequate T-cell response.
Overall, this study highlights the need for further research to understand the impact of SARS-CoV-2 on the immune system and to develop more effective treatments for COVID-19Discover how the team employed digital analysis of multiplex IF images to gain insights into the unique immune profile in fatal COVID-19 cases. Investigate the potential impacts of SARS-CoV-2 and related coronaviridae on antigen-presenting cells. Learn about the role played by the UKA in COVID-19 autopsies.
-
- Experience how the team used digital analysis of multiplex IF images to learn more about the specific immune landscape in fatal COVID cases
-
- Explore the possible effects of SARS-CoV-2 and related coronaviridae on antigen presenting cells
-
- Understand the role of the UKA in COVID-19 autopsies

Prof Ralf Huss, University Hospital Augsburg
Ralf Huss, MD, PhD, is a Professor of Pathology and the Managing Deputy Director of Pathology and Molecular Diagnostics at the University Hospital in Augsburg, Germany. He is also the head of the Center for Digital Medicine.
Huss holds board certifications in anatomical, experimental, and molecular pathology and boasts over 30 years of expertise in histopathology, immunology, cancer research, and oncology.

Lukas Borcherding, Medical Student, Technical University of Munich
Lukas Borcherding is a trained nurse who has been pursuing a degree in medicine at the Technical University of Munich, Germany since 2015. In 2020-2021, he took a hiatus from his studies to concentrate on the COVID-19 pandemic and examine the unique immune profile in fatal cases. Following the completion of his project, Lukas has resumed his studies and is presently finishing the practical component of his education. Upon receiving his license, he intends to work in internal medicine.