The immune response is spatially and temporally regulated. The density and location of immune cells in the tumor microenvironment (TME) have important diagnostic and prognostic values. Single cell-based multiomic technologies have exponentially increased our understanding of the numerous cellular and molecular networks regulating tumor initiation and progression. However, these techniques do not provide information about the spatial organization of cells or cell-cell interactions. Affordable, accessible, and easy to execute multiplexing techniques that allow spatial resolution of immune cells in tissue sections are needed to complement single cell-based high-throughput technologies.
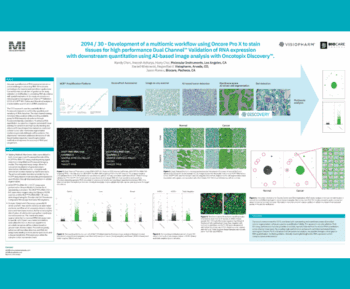
We have developed a strategy that integrates serial imaging, sequential labeling, and image alignment to generate virtual multiparameter slides of whole tissue sections. Virtual slides are subsequently analyzed in an automated fashion using the VIS software allowing us to identify, quantify, and map cell populations of interest. Specifically, the image analysis is performed using the analysis modules Tissuealign, Author, and HISTOmap. Here, we propose a strategy for the rational design of tissue multiplex assays using commercially available reagents, affordable microscopy equipment, and user-friendly software. Using this strategy, we created one virtual slide comprising 11 biomarkers plus two frequently used histological stains: hematoxylin and eosin (H&E) and picrosirius red (PSR). Multiple immune cell populations were identified, located, and quantified in different tissue compartments and their spatial distribution resolved using tissue heatmaps. This strategy maximizes the information that can be gained from limited clinical specimens and is applicable to formalin-fixed paraffin-embedded (FFPE) archived tissue samples, including whole tissue, core needle biopsies, and tissue microarrays. We propose this methodology as a useful guide for designing custom assays for identification, quantification, and mapping of immune cell populations in the TME.
Presented on March 22, 2021 at an XTalks webinar.
-
- Integration of serial imaging, sequential labeling, and image alignment in the experimental design of imaging assays can greatly increase the number of markers that can be visualized simultaneously, expand the possibilities of the analysis, and extract more information from precious clinical specimens.
-
- Virtual multiplexing allows to determine how markers visualized in one section spatially relate to markers visualized in another contiguous section.
-
- The use of whole tissue sections instead of selected fields of view for the analysis, results in an unbiased representation of the TME.
-
- The use of tissue heatmaps greatly simplify the visual representation of the spatial organization of cells in the tissue.

Manuel Flores, Ph.D. Candidate
Manuel Flores obtained his Biochemistry degree from Havana University, Cuba. In 2015 Manuel enrolled in the Immunology and Virology Master Program at Université de Montréal (Canada) and fast tracked to the Immunology and Virology PhD Program in 2016.
His doctoral research project focuses on characterizing the liver resident and infiltrating immune cell populations and their role in the pathogenesis of chronic liver diseases due to persistent viral and toxic injuries, including fibrosis and hepatocellular carcinoma. His research interests center around the spatial organization of immune cells in the hepatic tissue microenvironment, and the delineation of the multiple cell-cell interactions and their respective biological significance in health and disease. Manuel Flores is the recipient of doctoral scholarships from University of Montreal and from Fonds de recherche du Québec – Santé (FRQS).