Profiling fibroblastic heterogeneity: CAF functional states and their potential clinical applications in pancreatic cancer
Profiling fibroblastic heterogeneity: CAF functional states and their potential clinical applications in pancreatic cancer


Understanding the heterogeneity of the tumor microenvironment (TME) and its functional impact in solid tumors requires detailed examination of its cellular composition, the functional states of its cell populations, and their spatial organization within defined microenvironmental niches. This presentation will showcase data from an ongoing study utilizing the Harmonic Output of Stromal Traits (HOST), a research-driven framework for characterizing subsets of TME cells. HOST computes quantitative scores, called HOST-Factors, that distinguish pro-tumor and anti-tumor functional statuses across cellular populations and their respective niches. In particular, this talk will highlight the use of the HOST-Factor for profiling human pancreatic cancer-associated fibroblasts (CAFs) alongside key immunogenic and immunotolerant immune infiltrates, with a focus on evaluating their overall tumor-restrictive (TR) versus tumor-supportive (TS) roles.
Pancreatic ductal adenocarcinoma (PDAC) is profoundly influenced by its CAF-rich TME, which consists of active fibroblasts forming functional units with their self-generated extracellular matrix. To assess how this TME responds to therapy using the fibroblastic HOST-Factor, we are analyzing surgical specimens collected from a clinical trial at Fox Chase Cancer Center.
Our integrated workflow combines a curated set of CAF and immune cell biomarkers, automated high-plex immunofluorescence microscopy, and AI-guided image analysis using Visiopharm®. This pipeline assigns single-cell HOST-Factor values, enabling spatial mapping of functionally distinct CAFs and immune cells. By identifying TS and TR neighborhoods within the TME, this approach could offer novel insights into the fibroblastic and immune landscape’s response to therapy and open the possibility to support more precise patient stratification for personalized treatment decisions.

Janusz Franco-Barraza, MD, PhD
Janusz Franco-Barraza, MD, PhD is an Assistant Research Professor and Manager of the Spatial Immuno-Proteomics Facility at Fox Chase Cancer Center. His research is dedicated to uncovering how cancer-associated fibroblasts (CAFs) shape the tumor microenvironment and contribute to either tumor progression or restraint. Recognized for his expertise in high-plex immunofluorescence and AI-driven spatial analysis, Dr Franco-Barraza studies the dynamics of fibroblastic biology using relevant biomarker signatures that reveal the functional states of CAFs and their relationship to tumor development. His work provides insights that may help predict patient outcomes and responses to cancer therapies.
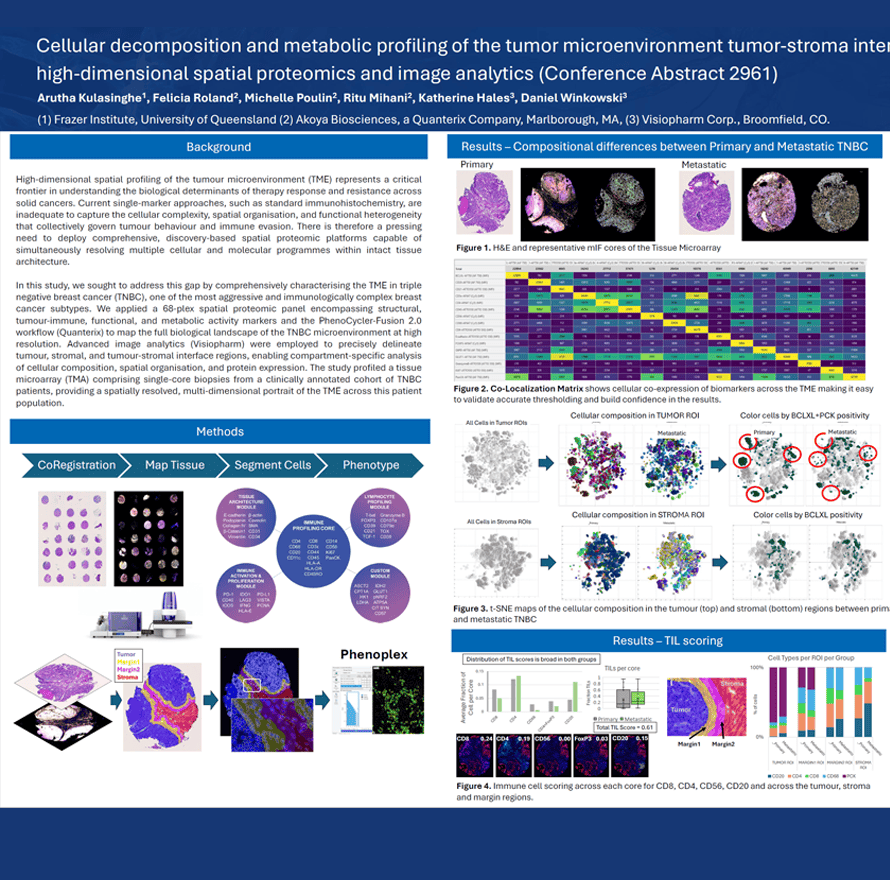

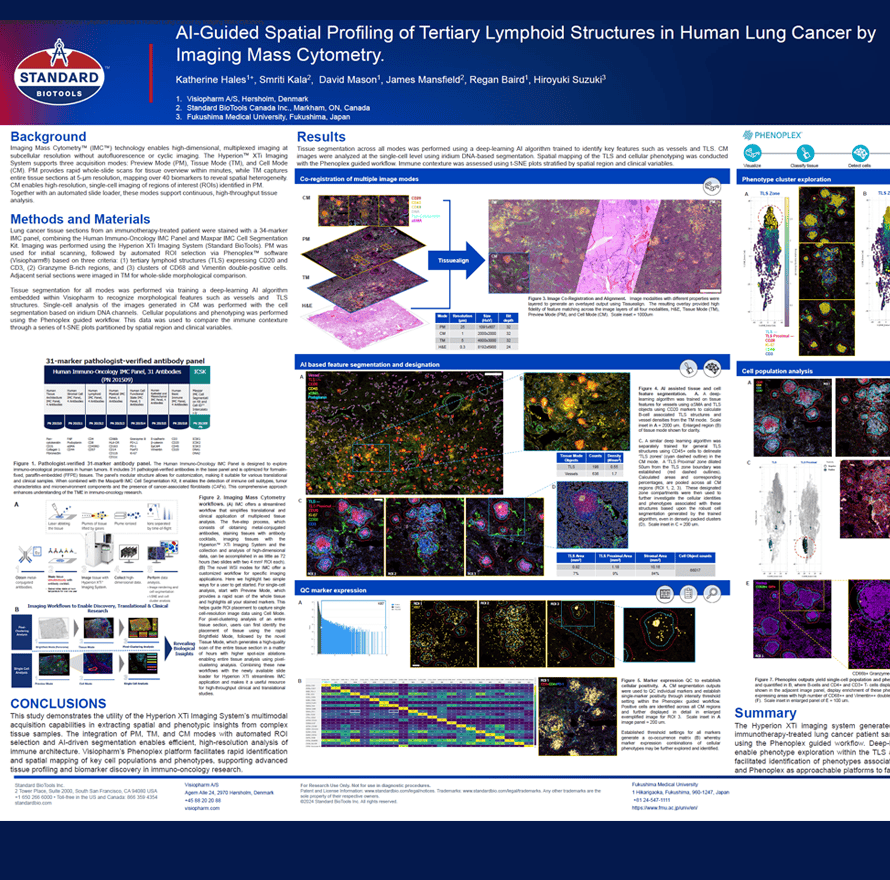
High-dimensional spatial profiling of the tumour microenvironment (TME) represents a critical frontier in understanding the biological determinants of therapy response and resistance across solid cancers. Current single-marker approaches, such as standard immunohistochemistry, are inadequate to capture the cellular complexity, spatial organisation, and functional heterogeneity that collectively govern tumour behaviour and immune evasion. There is therefore a pressing need to deploy comprehensive, discovery-based spatial proteomic platforms capable of simultaneously resolving multiple cellular and molecular programmes within intact tissue architecture.
In this study, we sought to address this gap by comprehensively characterising the TME in triple negative breast cancer (TNBC), one of the most aggressive and immunologically complex breast cancer subtypes. We applied a 68-plex spatial proteomic panel encompassing structural, tumour-immune, functional, and metabolic activity markers and the PhenoCycler-Fusion 2.0 workflow (Quanterix) to map the full biological landscape of the TNBC microenvironment at high resolution. Advanced image analytics (Visiopharm) were employed to precisely delineate tumour, stromal, and tumour-stromal interface regions, enabling compartment-specific analysis of cellular composition, spatial organisation, and protein expression. The study profiled a tissue microarray (TMA) comprising single-core biopsies from a clinically annotated cohort of TNBC patients, providing a spatially resolved, multi-dimensional portrait of the TME across this patient population.
Arutha Kulasinghe1, Felicia Roland2, Michelle Poulin2, Ritu Mihani2, Katherine Hales3, Daniel Winkowski, PhD3
- Frazer Institute, University of Queensland
- Akoya Biosciences, a Quanterix Company, Marlborough, MA
- Visiopharm Corp., Broomfield, CO.
 Sheila Hansen
Categories:
27964
Phenoplex™ – Overview
Sheila Hansen
Categories:
27964
Phenoplex™ – Overview
Phenoplex™ is Visiopharm’s end‑to‑end software solution for multiplex spatial biology analysis, enabling researchers to confidently visualize, segment, phenotype, and explore complex high‑plex tissue images within a single guided workflow. Built on AI‑powered cell detection and interactive verification steps, Phenoplex helps uncover meaningful biological insights, compare cohorts, and reproduce results across any plex level and imaging technology.
One platform.
Multiple modalities.
More answers.
The tumor microenvironment (TME) is a highly dynamic ecosystem that plays a central role in cancer progression, therapeutic resistance, and patient outcomes. However, its inherent complexity—driven by diverse immune, stromal, and malignant cell interactions—remains difficult to decode using conventional molecular profiling. Spatial proteomics technologies, such as Phenocycler Fusion/CODEX, now enable high-plex, protein-level mapping of cells within intact tissue architecture, providing unprecedented insights into both expression and localization patterns.
When coupled with artificial intelligence (AI), these approaches unlock a new dimension of discovery. AI methods, ranging from computer vision to graph-based modeling, can extract mechanistic insights, identify emergent spatial patterns, and uncover predictive signatures that are invisible to bulk or dissociated single-cell analyses. Our workflows integrate robust image preprocessing, segmentation, and spatial graph construction with interpretable AI platforms such as Visiopharm, enabling biologically meaningful target discovery.
Case studies illustrate how spatially resolved signatures can stratify patients for immunotherapy, reveal macrophage–tumor cell interactions and outperform conventional biomarkers in predicting clinical outcomes.
Despite significant challenges, such as batch effects, platform heterogeneity, and regulatory expectations for clinical deployment, a roadmap is emerging. Key elements include standardized panels and analysis pipelines, federated learning across multi-site cohorts, and integration of spatial biomarkers into companion diagnostics. By uniting computational, experimental, and clinical expertise, AI-driven spatial proteomics offers a direct path to more precise target selection, robust biomarker development, and improved patient stratification.
Ultimately, the convergence of spatial proteomics and AI has the potential to transform drug discovery and accelerate the development of next-generation cancer therapeutics that are both predictive and patient-centered.

Namrata Singh, PhD
Namrata Singh, PhD is a seasoned scientist and technical director at Dana-Farber Cancer Institute, with nearly three decades of experience spanning cancer research, spatial biology, and histological analysis. She also holds a concurrent research position at Harvard Medical School, where she contributes to cutting-edge immunohistochemistry and multiplex imaging studies.
Her expertise lies in high-dimensional tissue imaging and spatial analysis, leveraging advanced platforms such as Phenocycler Fusion, Vectra Polaris, Leica Bond RX, and analytical tools like Visiopharm, QuPath, and inForm. Namrata’s work bridges the gap between technology and translational research, enabling deeper insights into tissue architecture and disease pathology.
She earned her PhD and Master’s degrees from Delhi University, where she was a gold medalist.
Namrata is also an editor for research journals and holds numerous certifications in spatial biology technologies, including NanoString’s GeoMx DSP and nCounter platforms. Her contributions have been acknowledged internationally, including the second best paper presentation award from the IAEA.
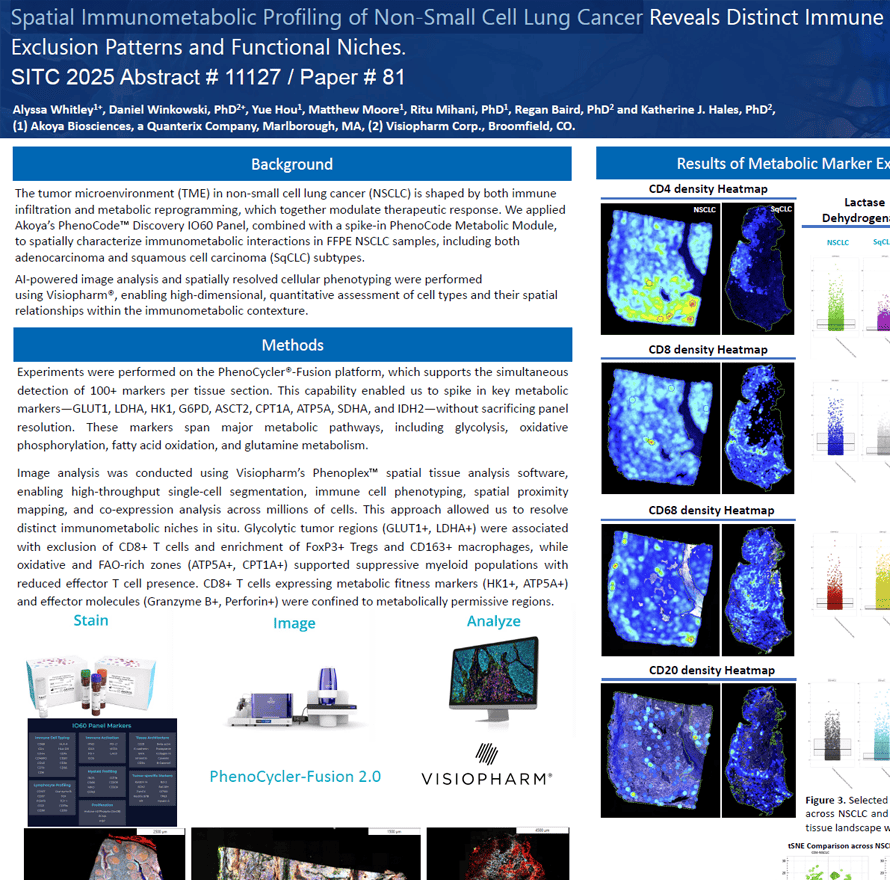

The tumor microenvironment (TME) in non-small cell lung cancer (NSCLC) is shaped by both immune infiltration and metabolic reprogramming, which together modulate therapeutic response. We applied Akoya’s PhenoCodeTM Discovery IO60 Panel, combined with a spike-in PhenoCode Metabolic Module, to spatially characterize immunometabolic interactions in FFPE NSCLC samples, including both adenocarcinoma and squamous cell carcinoma (SqCLC) subtypes.
AI-powered image analysis and spatially resolved cellular phenotyping were performed using Visiopharm®, enabling high-dimensional, quantitative assessment of cell types and their spatial relationships within the immunometabolic contexture.
Alyssa Whitley1+, Daniel Winkowski, PhD2+, Yue Hou1, Matthew Moore1, Ritu Mihani, PhD1, Regan Baird, PhD2 and Katherine J. Hales, PhD2
- Akoya Biosciences, a Quanterix Company, Marlborough, MA
- Visiopharm Corp., Broomfield, CO.
Imaging Mass CytometryTM (IMCTM) technology enables high-dimensional, multiplexed imaging at subcellular resolution without autofluorescence or cyclic imaging. The HyperionTM XTi Imaging System supports three acquisition modes: Preview Mode (PM), Tissue Mode (TM), and Cell Mode (CM). PM provides rapid whole-slide scans for tissue overview within minutes, while TM captures entire tissue sections at 5-μm resolution, mapping over 40 biomarkers to reveal spatial heterogeneity. CM enables high-resolution, single-cell imaging of regions of interest (ROIs) identified in PM. Together with an automated slide loader, these modes support continuous, high-throughput tissue analysis.
Katherine Hales1+, Smriti Kala2, David Mason1, James Mansfield2, Regan Baird1, Hiroyuki Suzuki3
- Visiopharm A/S, Hørsholm, Denmark
- Standard BioTools Canada Inc., Markham, ON, Canada
- Fukushima Medical University, Fukushima, Japan