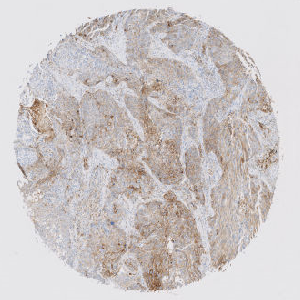
Overview of a single PD-L1 stained TMA core with melanoma tissue.
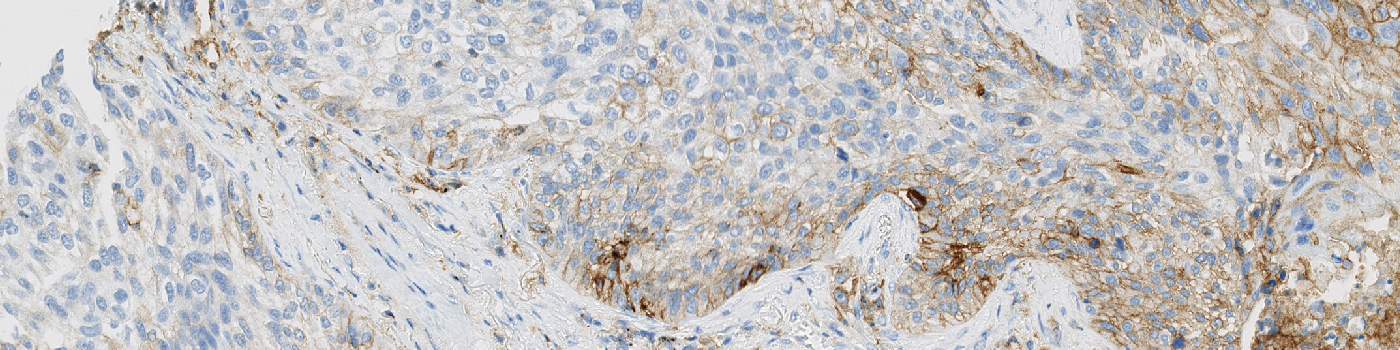
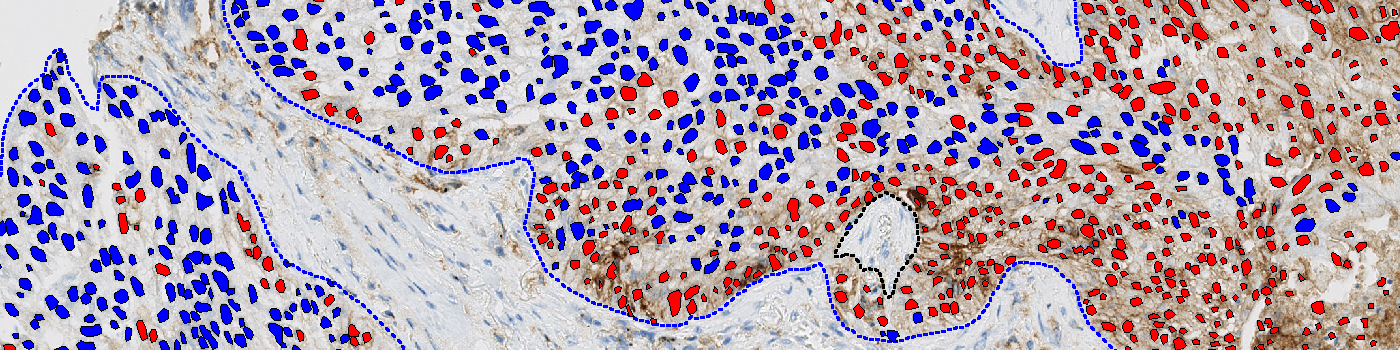
#10123
Programmed death-ligand 1 (PD-L1) is a transmembrane protein that binds to the inhibitory receptor programmed death 1 receptor (PD-1), causing a down regulation of immune responses. PD-L1 is typically expressed on normal cells but has been observed in immune cells and tumor cells, see [1], while PD-1 is typically expressed on cytotoxic T-cells and other immune cells. Tumor cells can upregulate PD-L1 expression and avoid being attacked by the body’s immune system, making an interruption of the PD-1/PD-L1 interaction an attractive method for assisting the immune system in destroying tumor cells, see [2].
This APP can be used to determine the proportion of PD-L1 positive cells. It quantifies all cells and classifies PD-L1 positive cells based on brown/DAB positive membrane staining.
Auxiliary APPs
APP: 01 Tumor Detect
The Tumor Detection APP is used for initial tumor region detection and is included with the APP.
Quantitative Output variables
The output variables obtained from this protocol are:
Workflow
Step 1: Use the Tumor Detect APP and/or outline tumor regions manually if necessary.
Step 2: Run the PD-L1 APP. Click the save button to transfer the results to the database.
Methods
A hematoxylin input band is filtered with a median filter and combined with image intensity to smoothen out and mitigate granularity of nuclei. Nuclei are then detected using two different size scales of polynomial blob filters in combination with polynomial local linear filters that delimit the cells. Membranes are detected using two different scales of polynomial local linear filters on a DAB input band. Postprocessing steps grow the nuclei blobs and combine/separate adjacent nuclei appropriately to match the hematoxylin band. The classified PD-L1 positive membrane staining is verified by size, shape and intensity constraints and then applied to the adjacent cells to quantify the number of PD-L1 positive cells.
Staining Protocol
Staining protocols have been developed by Dako and Ventana for the following three assays which have been found to produce comparable results, see [3], and are available from their website(s):
Keywords
PD-L1, PD-1, T-cell, cancer, oncology, IHC, membrane, skin, melanoma, digital pathology, image analysis, B7-H1, hematoxylin
References
USERS
This APP was developed in collaboration with NordiQC.
LITERATURE
1. Zou, W., Chen, L. Inhibitory B7-family molecules in the tumour microenvironment, Nat Rev Immunology 2008, 8, 467-477, DOI.
2. Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy, Nat Rev Cancer 2012, 12 (4), 252-264, DOI.
3. Hirsch, F. R., et. al. PD-L1 immunohistochemistry assays for lung cancer: results from phase 1 of the blueprint PD-L1 IHC assay comparison project, Journal of Thoracic Oncology 2017, 12 (2), 208-222, DOI.