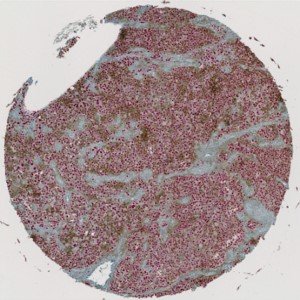
Overview of a single PD-L1+SOX10 stained TMA core with melanoma tissue.
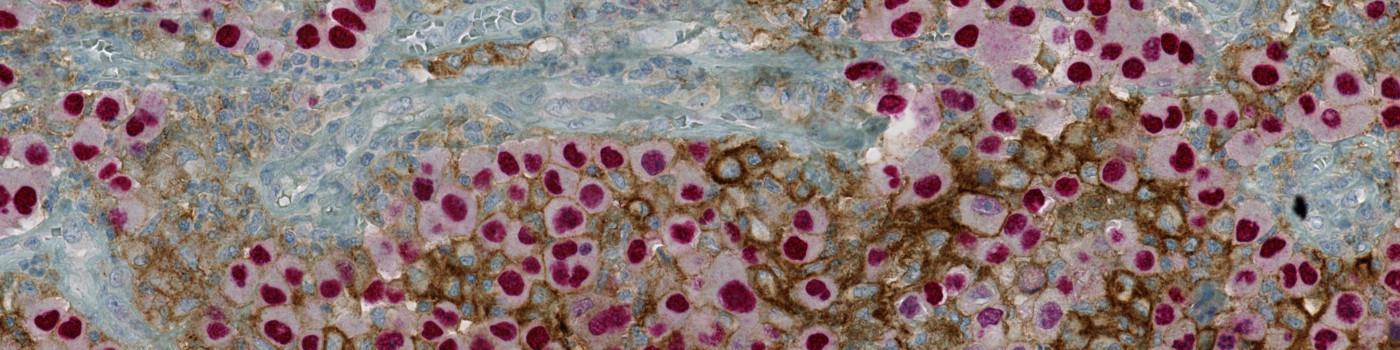
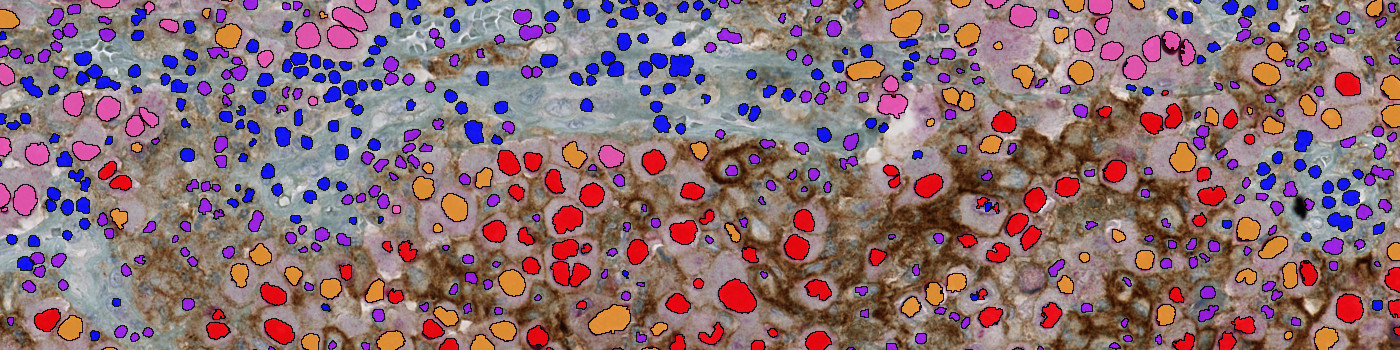
#10125
Programmed death-ligand 1 (PD-L1) is a transmembrane protein that binds to the inhibitory receptor programmed death 1 receptor (PD-1), causing a down regulation of immune responses. PD-L1 is typically expressed on normal cells but has been observed in immune cells and tumor cells, see [1]. while PD-1 is typically expressed on cytotoxic T-cells and other immune cells. Tumor cells can upregulate PD-L1 expression and avoid being attacked by the body’s immune system, making an interruption of the PD-1/PD-L1 interaction an attractive method for assisting the immune system in destroying tumor cells, see [2].
SOX10 is a nuclear transcription factor that has been shown to be a sensitive and specific marker for malignant melanoma, see [3]. Combining SOX10 and PD-L1 enables the user to assess whether PD-L1 expression originates from tumor cells or adjacent non-tumor cells.
This APP can be used to determine the number of tumor and non-tumor cells that express PD-L1 without having to outline tumor regions.
Quantitative Output variables
The output variables obtained from this protocol are:
Workflow
Run the APP. Click the save button to transfer the results to the database.
Methods
Based on an AEC red enhanced input band, SOX10 positive nuclei are classified using a combination of polynomial blob- and local linear filters. SOX10 negative nuclei are classified in the same manner but using a hematoxylin input band. Membranes are detected using two different scales of polynomial local linear filters on green/blue contrast and brown chromogen input bands. Postprocessing steps grow the nuclei blobs and combine/separate adjacent nuclei appropriately to match the input band. The classified PD-L1 positive membrane staining is verified by size, shape and intensity constraints and then applied to the adjacent cells to quantify the number of PD-L1 positive cells. An algorithm evaluates PD-L1 expression shared between SOX10 negative and positive cells to determine if tumor cells are truly PD-L1 positive or influenced by adjacent non-tumor cells.
Staining Protocol
There is no staining protocol available.
Keywords
PD-L1, PD-1, T-cell, cancer, oncology, IHC, membrane, skin, melanoma, digital pathology, image analysis, B7-H1, SOX10, hematoxylin
References
USERS
This APP was developed in collaboration with Aalborg University Hospital.
LITERATURE
1. Zou, W., Chen, L. Inhibitory B7-family molecules in the tumour microenvironment, Nature Reviews Immunology 2008, 8 (6), 467-477, DOI.
2. Pardoll, D. M.The blockade of immune checkpoints in cancer immunotherapy, Nat Rev Cancer 2012, 12 (4), 252-264, DOI.
3. Efternavn, Fornavnsinitialer (evt. et. al.) SOX10: a useful marker for identifying metastatic melanoma in sentinel lymph nodes, Applied Immunohistochemistry & Molecular Morphology 2015, 23 (2), 109-112, DOI.