AI & spatial proteomics: unlocking the TME for next-gen cancer drugs: A new paradigm in cancer drug discovery through spatial intelligence


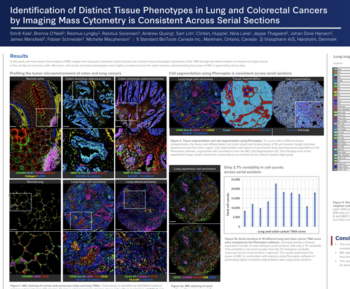
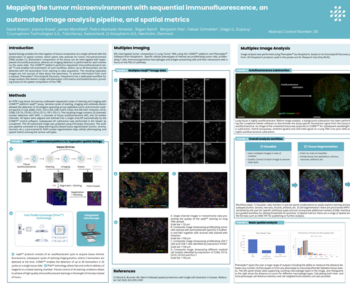
The tumor microenvironment (TME) is a highly dynamic ecosystem that plays a central role in cancer progression, therapeutic resistance, and patient outcomes. However, its inherent complexity—driven by diverse immune, stromal, and malignant cell interactions—remains difficult to decode using conventional molecular profiling. Spatial proteomics technologies, such as Phenocycler Fusion/CODEX, now enable high-plex, protein-level mapping of cells within intact tissue architecture, providing unprecedented insights into both expression and localization patterns.
When coupled with artificial intelligence (AI), these approaches unlock a new dimension of discovery. AI methods, ranging from computer vision to graph-based modeling, can extract mechanistic insights, identify emergent spatial patterns, and uncover predictive signatures that are invisible to bulk or dissociated single-cell analyses. Our workflows integrate robust image preprocessing, segmentation, and spatial graph construction with interpretable AI platforms such as Visiopharm, enabling biologically meaningful target discovery.
Case studies illustrate how spatially resolved signatures can stratify patients for immunotherapy, reveal macrophage–tumor cell interactions and outperform conventional biomarkers in predicting clinical outcomes.
Despite significant challenges, such as batch effects, platform heterogeneity, and regulatory expectations for clinical deployment, a roadmap is emerging. Key elements include standardized panels and analysis pipelines, federated learning across multi-site cohorts, and integration of spatial biomarkers into companion diagnostics. By uniting computational, experimental, and clinical expertise, AI-driven spatial proteomics offers a direct path to more precise target selection, robust biomarker development, and improved patient stratification.
Ultimately, the convergence of spatial proteomics and AI has the potential to transform drug discovery and accelerate the development of next-generation cancer therapeutics that are both predictive and patient-centered.

Namrata Singh, PhD
Namrata Singh, PhD is a seasoned scientist and technical director at Dana-Farber Cancer Institute, with nearly three decades of experience spanning cancer research, spatial biology, and histological analysis. She also holds a concurrent research position at Harvard Medical School, where she contributes to cutting-edge immunohistochemistry and multiplex imaging studies.
Her expertise lies in high-dimensional tissue imaging and spatial analysis, leveraging advanced platforms such as Phenocycler Fusion, Vectra Polaris, Leica Bond RX, and analytical tools like Visiopharm, QuPath, and inForm. Namrata’s work bridges the gap between technology and translational research, enabling deeper insights into tissue architecture and disease pathology.
She earned her PhD and Master’s degrees from Delhi University, where she was a gold medalist.
Namrata is also an editor for research journals and holds numerous certifications in spatial biology technologies, including NanoString’s GeoMx DSP and nCounter platforms. Her contributions have been acknowledged internationally, including the second best paper presentation award from the IAEA.